We are a molecular genetics lab that studies the function of NUCLEAR FACTOR Y (NF-Y) transcription factors in the plant lineage.
Nuclear Factor Y (NF-Y) Transcription Factors
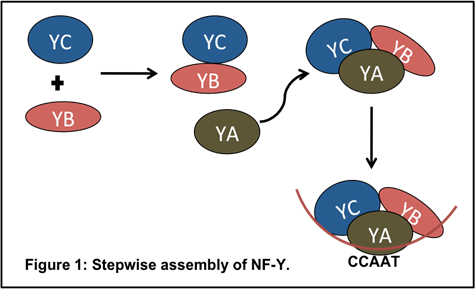
NF-Y transcription factors are composed of three independent protein families, NF-YA, NF-YB, and NF-YC. To activate target genes, NF-YB and NF-YC dimerize in the cytoplasm and move to the nucleus where the heterodimer interacts with NF-YA to create the DNA-binding, heterotrimeric NF-Y transcription factor complex.
NF-Y transcription factors are composed of three independent protein families, NF-YA, NF-YB, and NF-YC. To activate target genes, NF-YB and NF-YC dimerize in the cytoplasm and move to the nucleus where the heterodimer interacts with NF-YA to create the DNA-binding, heterotrimeric NF-Y transcription factor complex.
The NF-Y is ubiquitous in eukaryotes; in animals, the NF-Y plays a crucial role in cell cycle regulation and is an interactor of the transcription factor p53, a key protein involved in tumor suppression. In plants, NF-Y is primarily known for the role in regulating photoperiod-dependent flowering.
Network analysis of the NF-Y transcription factors
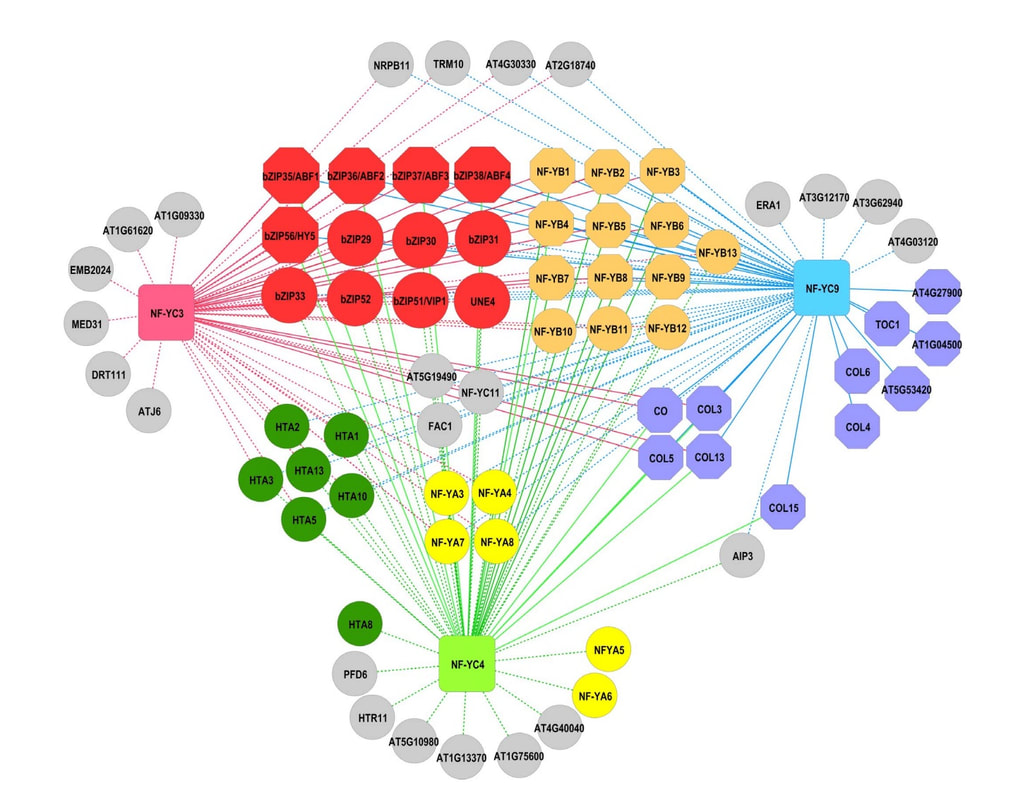
Network analysis is a type of bioinformatic analysis that visualizes complex relationships between genes and/or proteins in a biological process. Network analysis is widely used to predict gene function and identify novel components of a molecular pathway. The findings of the bioinformatic analysis are then confirmed by wet-lab experiments that employ molecular tools such as gene cloning, real-time PCR (gene coexpression), and Yest 2-hybrid (protein-protein interactions).
Our research uses network analysis to elucidate the function of the NF-Y in plant systems. The bioinformatic analysis is followed by wet-lab experiments that validate key interactions that are identified. The wet lab experiments we perform include, DNA extraction, gene cloning, polymerase chain reaction, agarose gel electrophoresis, Yeast 2-hybrid, protein extraction, SDS -PAGE electrophoresis, Western blotting, RNA extraction, and quantitative real-time PCR.
Network analysis is a type of bioinformatic analysis that visualizes complex relationships between genes and/or proteins in a biological process. Network analysis is widely used to predict gene function and identify novel components of a molecular pathway. The findings of the bioinformatic analysis are then confirmed by wet-lab experiments that employ molecular tools such as gene cloning, real-time PCR (gene coexpression), and Yest 2-hybrid (protein-protein interactions).
Our research uses network analysis to elucidate the function of the NF-Y in plant systems. The bioinformatic analysis is followed by wet-lab experiments that validate key interactions that are identified. The wet lab experiments we perform include, DNA extraction, gene cloning, polymerase chain reaction, agarose gel electrophoresis, Yeast 2-hybrid, protein extraction, SDS -PAGE electrophoresis, Western blotting, RNA extraction, and quantitative real-time PCR.